In 1907, a young Ross Granville Harrison published observations on the outgrowth of embryonic frog neurons1. Little did he know his work would put him in the running for a Nobel prize2. At the time, the mechanics of neuronal outgrowth were poorly understood due in part to the complexity of the developing nervous system. Harrison reasoned that definitive answers could only be found by removing neurons from the embryo and studying them in isolation. Some tissues had been kept alive outside of the body for short periods of time, but no one had grown individual cells outside of a living organism. No one, that is, until Harrison.
Now, more than a century later, Harrison’s work is viewed as the origin of modern two-dimensional (2D) cell culture—a foundational approach to researching biological processes. By isolating cells from the dynamic environment of a living organism, researchers can gather high-resolution observations of a cell’s behavior and what influences it. Using 2D culture, scientists have discovered vaccines for polio3, elucidated mechanisms of cancer biology4, and much more.
Though beneficial, 2D cell culture suffers from its artificial nature5,6. Cells grown in isolation, atop plastic dishes, in stagnant media, behave abnormally relative to cells in vivo. As such, the results gained in 2D cell culture may not accurately predict cell behavior in vivo.
To improve the accuracy of cell culture while preserving its many benefits, researchers have developed three-dimensional (3D) culture techniques that encourage in vivo-like cell behavior, enabling them to more accurately model human physiology, disease, and response to therapeutics. Read on for a brief overview of what 3D cell culture is and how it is advancing in vitro studies.
2D cell culture: A valuable, yet flawed, foundation
True to Harrison’s rationale, 2D cell culture remains a powerful method for studying cell and molecular biology. Typically, 2D cell culture refers to the growth or maintenance of cells in a single monolayer on top of glass or plastic dishes, where a nutrient-rich media nourishes the cells and provides a livable environment.
Some of the benefits of 2D cell culture include:
- Accessibility of high-resolution data: Observing the inner workings of a cell when it’s tucked away inside a living organism can be extremely challenging. But in 2D culture, there are no dense layers of tissue and extracellular proteins to scatter light and distort sound. Under these conditions, various techniques can be used to collect high-resolution observations of a cell’s behavior and inner workings.
- Ease of use: Researchers can quickly become proficient with 2D cell culture, as it is relatively non-technical, requires very little space, and consumes inexpensive reagents.
- High throughput: Since 2D cell culture is easy and inexpensive, researchers can build automated, highly parallelized workflows that enable large-scale, high-throughput experimentation.
- Well-established: Scientists have been working with 2D cell culture for over a century, providing a large collection of historical data to compare to for quality control and performance metrics.
Though common and beneficial in many ways, 2D cell culture is highly artificial. In Harrison’s own observations, he noted that the abnormal growth of cells in vitro “might be expected even on purely mechanical grounds, for the stresses and strains which are brought to bear upon the [cells in vitro] must be entirely different from those within the intact embryo.”1
Harrison is pointing out that cells grown under 2D conditions are exposed to a very different environment than those in vivo.
These disadvantages of 2D cell culture include:
- Lack of cellular diversity: 2D culture is often performed with only one or two cell types resulting in limited cell-cell interactions, which critically influence cell behavior in vivo.
- Reliance on non-physiological cell types: The static microenvironment limits the ability to culture physiologically relevant primary human or iPS-derived cells, often limiting cell culture to cancerous or immortalized cell lines that do not behave like healthy cells in the human body.
- Lack of nutrient, oxygen, and waste gradients: In vivo, all cells are within a few hundred micrometers of blood vessels or perfusing liquids, placing them in overlapping oxygen, sugar, nutrient, and cell waste gradients. The cells position along these gradients affects its gene expression and behavior. Cells grown in 2D cell culture are exposed to very few gradients and are often grown in abnormally high oxygen and sugar concentrations.
- Lack of mechanical input: The stagnant 2D environment lacks natural mechanical cues, such as fluid flow or periodic contractions, both of which are known to influence cellular behavior and disease.
- Impaired polarity: When grown in a monolayer, the combination of a rigid, impenetrable substrate below the cells with the absence of cells above them can lead to an abnormal or suboptimal polarization.
Collectively, these factors cause abnormal behavior and decrease the likelihood that results gathered in 2D cell cultures will reflect in vivo results. Given these drawbacks, researchers have long been interested in developing culture methods that encourage cells to behave naturally, leading to the creation of 3D cell culture systems.
3D cell culture: A more accurate model of in vivo conditions
3D cell culture systems grow cells in a more human-relevant microenvironment.
This results in several advantages of 3D cell culture, including:
- Natural gradient formation: Under 3D culture, nutrients, oxygen, drugs, and other diffuse materials will naturally form gradients due to the selective permeability of this environment. The recapitulation of these gradients provides researchers with unique insights into drug penetration and spatially dependent cell behaviors that might occur in the human body.
- Increased cellular diversity: 3D cultures often include heterogeneous cell populations, enabling the recreation of critical natural cell-cell interactions when modeling inflammation, angiogenesis, and other physiological processes.
- Physiologically relevant mechanical forces: Advanced 3D culture systems mimic the biomechanical forces cells experience in vivo, including fluid flow and cyclic stretching (to emulate in vitro fluid flow and breathing or peristalsis, respectively).
- In vivo-like cell morphology and gene expression: Collectively, the above factors encourage cells to exhibit in vivo-like gene expression profiles, polarities, and morphologies.
Types of 3D models
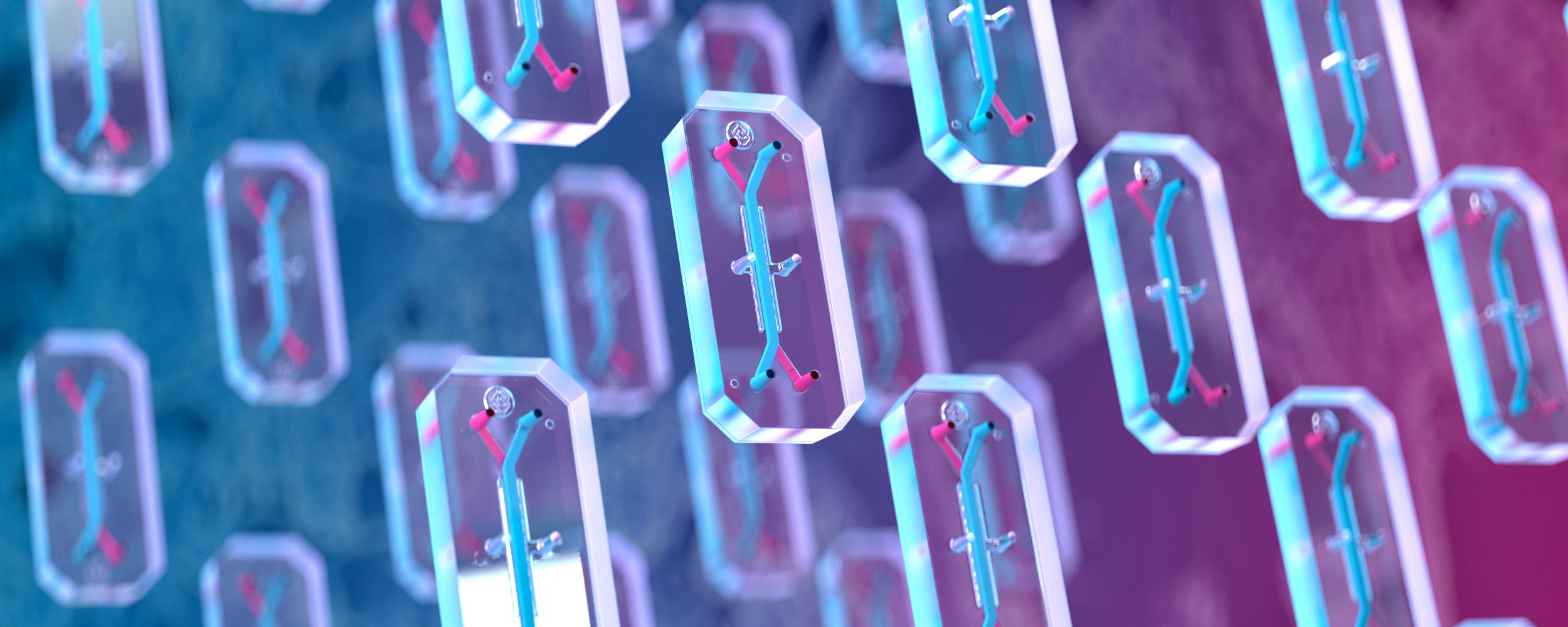
The two most common types of 3D cell culture systems are spherical culture models and microphysiological systems like Organ-Chips5,6.
Spherical culture models encompass various types of spheroid and organoid models that culture cells in a 3D mass on top of an inert substrate or on extracellular matrix (ECM) proteins, as well as those which embed cells in ECM proteins. These models are frequently used to study stem cell niches and to model the tumor microenvironment5,6,7.
While a step up from 2D cell culture, spherical models are limited by their lack of both perfusion and mechanical cues as well as their abnormal architecture. This is where Organ-Chips excel.
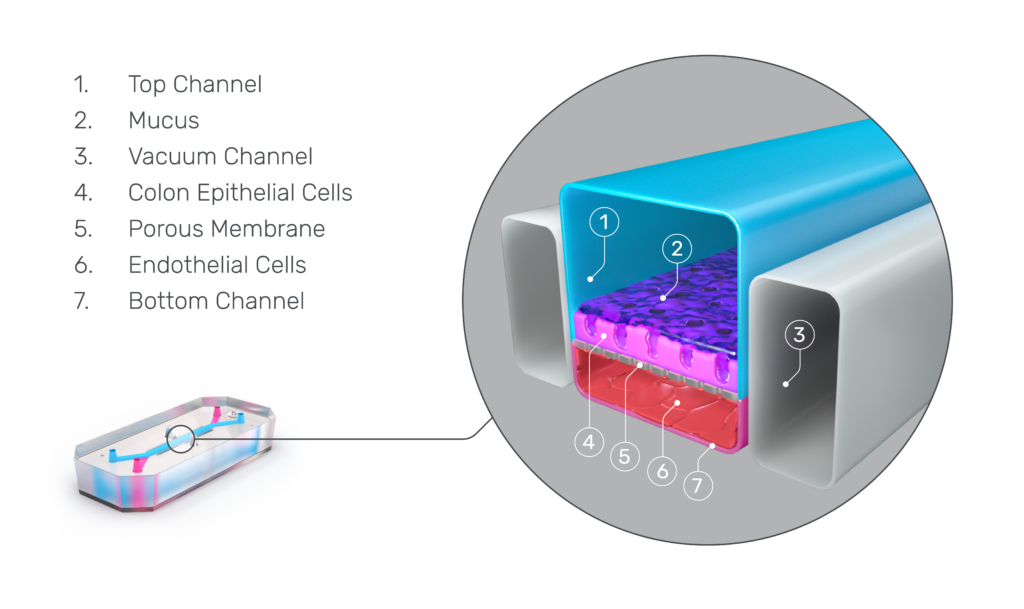
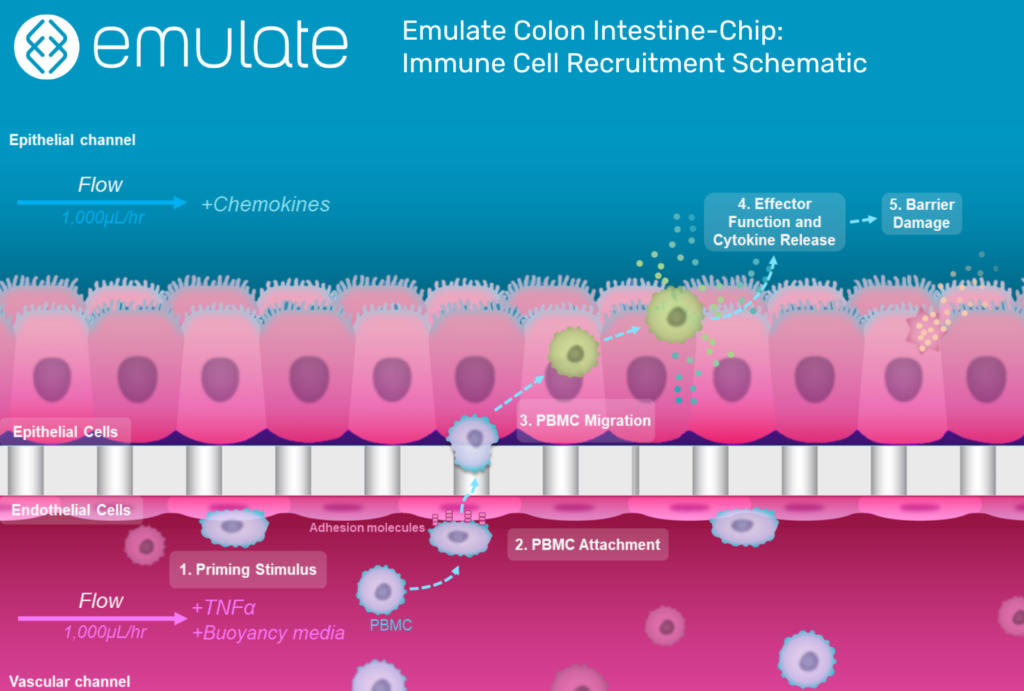
Organ-Chips are more complex, microphysiological systems that combine tissue-specific ECM proteins, heterogeneous cell populations, and mechanical cues in a structured framework that organizes cells to recapitulate tissue-level architecture. Among 3D models, Organ-Chips are the most realistic model of in vivo physiology and have been used to study numerous organs and diseases8.
The key advantages of Organ-Chips over spherical models include:
- Media perfusion: Organ-Chips replicate natural fluid flow found in vivo, preventing waste buildup and establishing nutrient and oxygen gradients.
- Higher-level organization: By seeding cells in a structured framework, Organ-Chips produce organized cell layers and barriers, facilitating more in vivo-like cell-cell interactions.
- Mechanical forces: Organ-Chips can be compressed and relaxed to mimic breathing or peristalsis. This, along with the shear forces generated by fluid flow in the chips, influences cell movement and gene expression.
Given these benefits and the fact that Organ-Chips incorporate real human cells, it’s not surprising that they’re increasing in popularity—particularly in the drug development space.
Improving drug development with Organ-Chips
Predicting how compounds will behave in the complex environment of the human body has been a persistent challenge in drug development. Since they are themselves living systems, animals have long stood as the go-to model for pharmaceutical developers to test how drugs may act in humans before clinical trials. However, species differences are great enough that the safety and efficacy of drug candidates seen in animals often fail to translate to human response.
Organ-Chips are a valuable tool in drug development because they enable researchers to test how drugs affect human cells by using human cells. Unlike 2D cell culture, Organ-Chips can be used to see how drugs diffuse across tissue barriers, how different cell types react to the drug, and how these reactions affect tissue integrity.
It has been more than a century since Harrison performed the first in vitro cell culture experiment. While 2D cell culture remains a critical component of modern research, technological advancements have led to an evolution in cell culture towards more human-relevant 3D models—and Emulate has been at the forefront of this evolution. Researchers from Emulate have developed numerous Organ-Chips (including models of the brain, liver, kidney, intestine, and more), that are already being used by researchers across pharma and academia to study complex biological processes in unprecedented detail, accelerating the development of safer, more effective therapeutics to improve human health.
- Harrison, Rose G., et al. “Observations of the Living Developing Nerve Fiber.” The Anatomical Record, vol. 1, no. 5, 1 June 1907, pp. 116–128, 10.1002/ar.1090010503.
- “Yale Alumni Magazine: America’s Most Famous Unknown Scientist (Feb 02).” Web.archive.org, 14 Nov. 2012, web.archive.org/web/20121114035855/yalealumnimagazine.com/issues/02_02/old_yale.html. Accessed 13 May 2022.
- Turner, Timothy. “Development of the Polio Vaccine: A Historical Perspective of Tuskegee University’s Role in Mass Production and Distribution of HeLa Cells.” Journal of Health Care for the Poor and Underserved, vol. 23, no. 4a, 2012, pp. 5–10, www.ncbi.nlm.nih.gov/pmc/articles/PMC4458465/, 10.1353/hpu.2012.0151.
- Davies, Preston. “Significant Research Advances Enabled by HeLa Cells – Office of Science Policy.” Office of Science Policy, 2010, osp.od.nih.gov/scientific-sharing/hela-cells-timeline/.
- Jensen, Caleb, and Yong Teng. “Is It Time to Start Transitioning from 2D to 3D Cell Culture?” Frontiers in Molecular Biosciences, vol. 7, 2020, p. 33, www.ncbi.nlm.nih.gov/pubmed/32211418, 10.3389/fmolb.2020.00033.
- Research, Center for Drug Evaluation and. “Three-Dimensional (3D) Cell Culture (Microphysiological) Platforms as Drug Development Tools.” FDA, 23 Aug. 2021, www.fda.gov/drugs/regulatory-science-action/three-dimensional-3d-cell-culture-microphysiological-platforms-drug-development-tools. Accessed 13 May 2022.
- Riffle, Stephen, and Rashmi S. Hegde. “Modeling Tumor Cell Adaptations to Hypoxia in Multicellular Tumor Spheroids.” Journal of Experimental & Clinical Cancer Research, vol. 36, no. 1, 3 Aug. 2017, 10.1186/s13046-017-0570-9. Accessed 9 Apr. 2019.
- Ingber, Donald E. “Human Organs-On-Chips for Disease Modelling, Drug Development and Personalized Medicine.” Nature Reviews Genetics, 25 Mar. 2022, pp. 1–25, www.nature.com/articles/s41576-022-00466-9, 10.1038/s41576-022-00466-9. Accessed 9 May 2022.
- Ewart, Lorna, et al. Qualifying a Human Liver-Chip for Predictive Toxicology: Performance Assessment and Economic Implications. 16 Dec. 2021, 10.1101/2021.12.14.472674.