Earlier this week, The New York Times ran an article outlining the need to build a strategic monkey reserve to address shortages in rhesus monkeys that have arisen in the testing of COVID-19 vaccines. Shortages have arisen since the import of animals from China have been banned during the pandemic, causing prices of monkeys to skyrocket. But the article fails to address the real issue—it’s time to say, ‘enough is enough’ and begin to discourage the use of primates for drug testing. Our societal goal should be reducing the number of monkeys used and encouraging more innovative alternatives to primate testing, not breeding more laboratory animals to keep up with the high demand from drug developers.
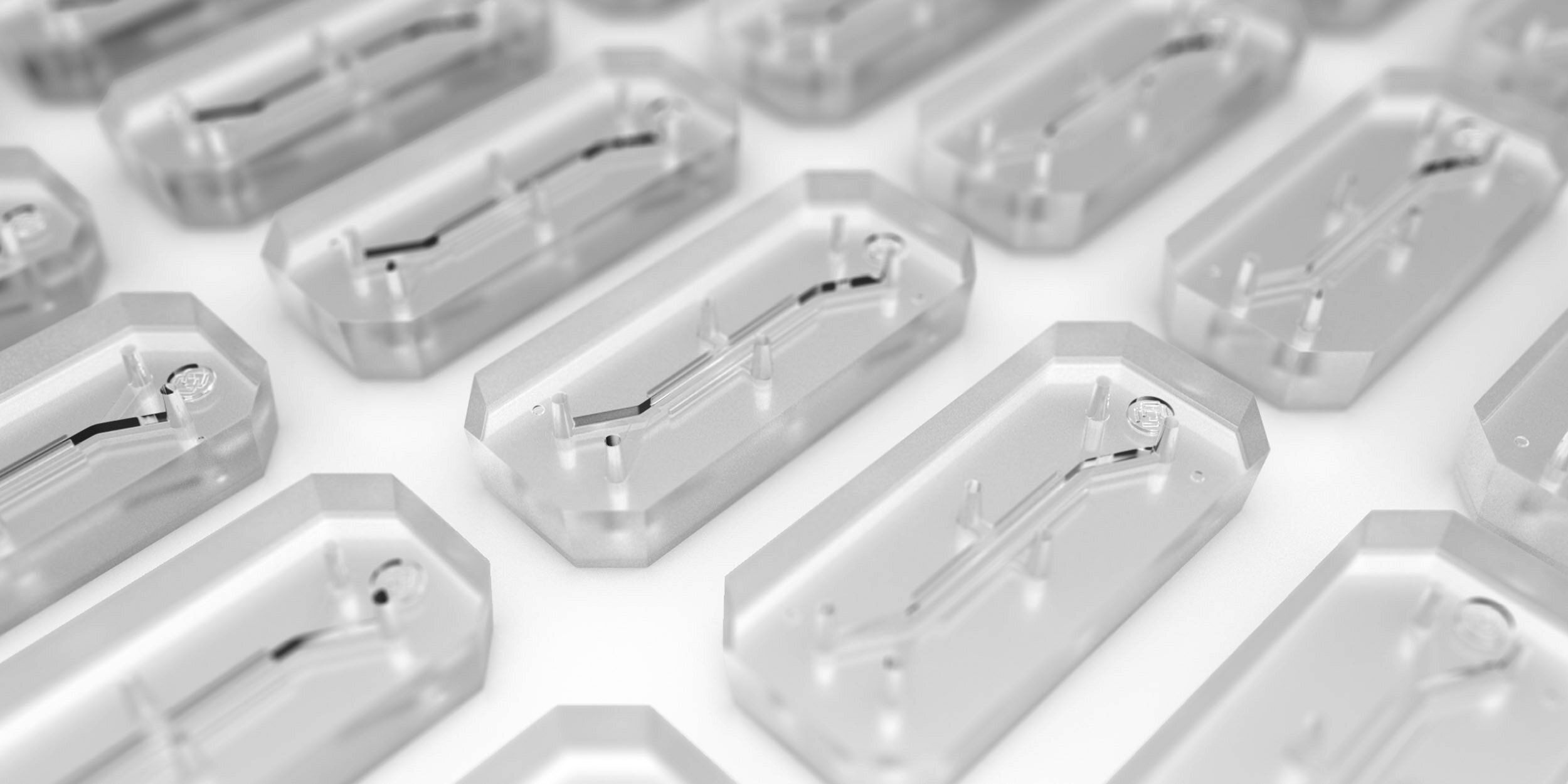
Beyond the obvious ethical concerns, animal testing has severe limitations in effectiveness, particularly for today’s novel biologic drugs and vaccines. Monkeys are not always capable of replicating human disease processes or drug response. Advances in biopharmaceutical research, including Organ-on-a-Chip technology, which recreates human biology in highly controlled lab settings, can more closely emulate human drug response.
While animal testing may always play a role in some pharmaceutical testing, a strategic monkey reserve is not the answer. It’s our responsibility to explore alternatives. If better patient outcomes are the end game, we should encourage the use of technology that gets us there faster, and more humanely. We are, however, starting to see progress from a policy perspective in recognizing the importance of new alternatives to primate (and other animals) testing. In 2020, Congressional Representatives Alcee L. Hastings (D-Florida) and Vern Buchanan (R-Florida) together introduced legislation that would help improve animal testing transparency, while incentivizing the study of more human-relevant testing methods. We hope that similar legislation is introduced into the new Congress, as we believe it is the first step towards replacing animal testing with new methods that can more effectively recreate human cellular responses.